The new tool brings together cutting-edge AI models under one roof, saving scientists precious time they'd otherwise spend wrestling with code. DeepProtein tackles everything from predicting how proteins fold to mapping their interactions with other molecules. It's built for both AI experts and biologists who just want their protein analysis to work without a PhD in computer science.
The team didn't just build a tool – they put it through its paces. They tested eight different types of AI architectures across multiple protein analysis tasks. These ranged from basic classification problems to the more complex challenge of predicting protein structures in 3D space.
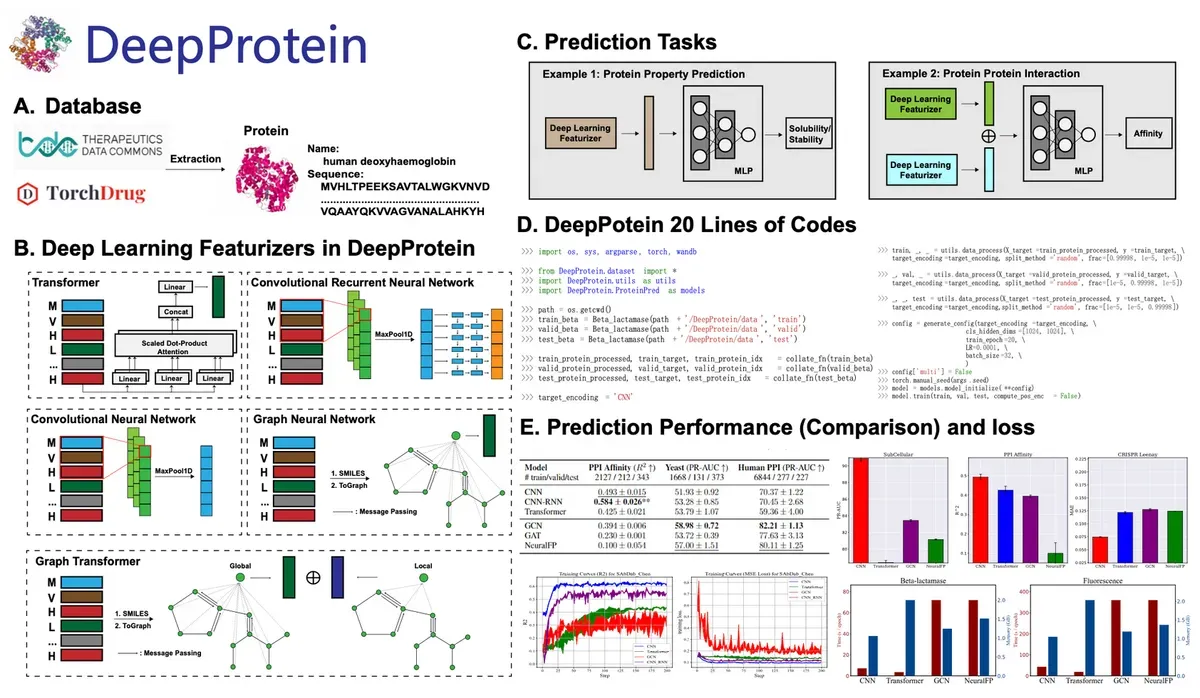
The star of the show is their new model family, DeepProt-T5. Based on the powerful Prot-T5 architecture, these fine-tuned models achieved top scores on four benchmark tasks and strong results on six others. Think of it as a straight-A student who also plays varsity sports.
What sets DeepProtein apart is its user-friendly approach. Previous tools often required researchers to understand both complex biology and deep learning. DeepProtein strips away this complexity with a simple command-line interface. It's like having an AI research assistant who speaks plain English.
The library builds on DeepPurpose, a widely used tool for drug discovery. This heritage means researchers can easily integrate DeepProtein with existing workflows and databases. The team also provides detailed documentation and tutorials, ensuring scientists don't get stuck in implementation details.
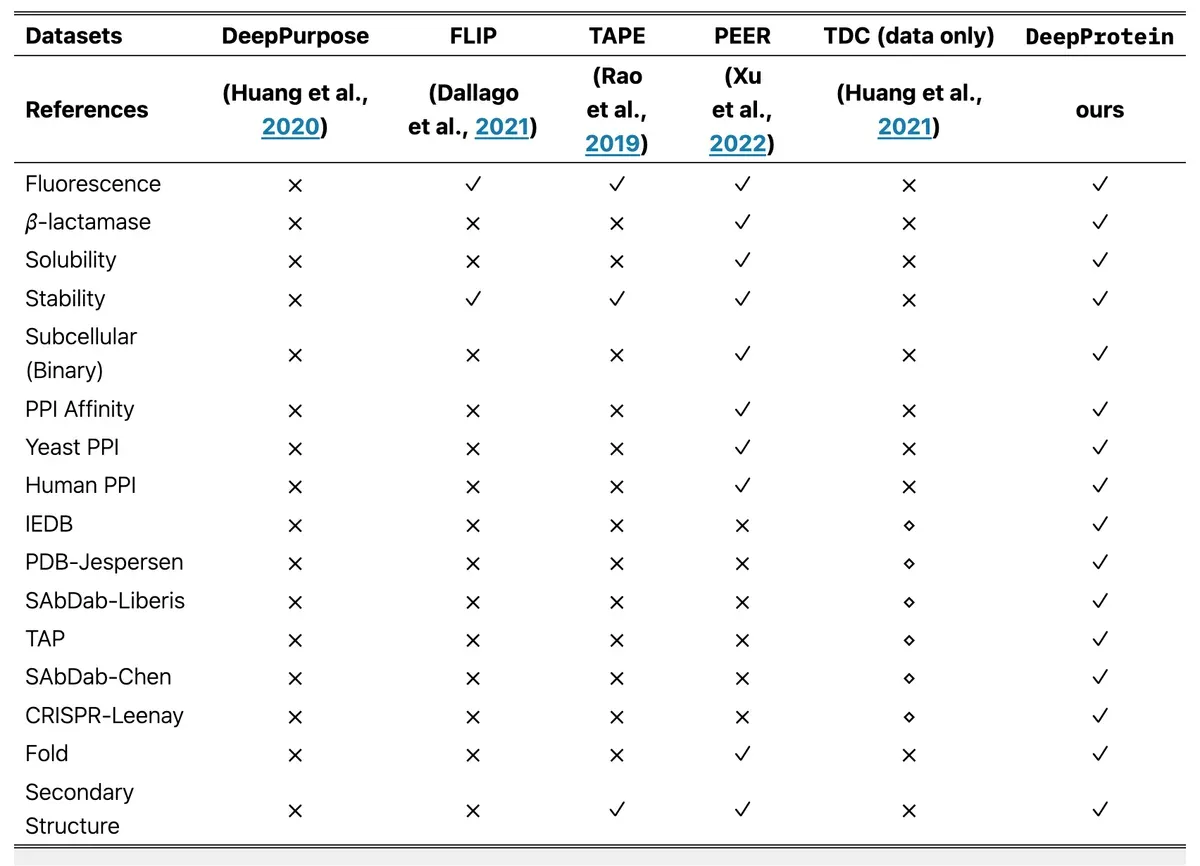
DeepProtein fills several gaps in the protein research toolkit. While previous benchmarks like PEER focused mainly on sequence-based methods, DeepProtein adds structure-based approaches and pre-trained language models to the mix. It's the difference between having just a hammer and owning a complete toolbox.
The timing couldn't be better. The success of tools like AlphaFold 2.0 has sparked renewed interest in applying machine learning to protein research. DeepProtein rides this wave by making advanced AI techniques accessible to more researchers.
For the technically minded, the library supports various neural network architectures: CNNs, CNN-RNNs, RNNs, transformers, graph neural networks, graph transformers, pre-trained protein language models, and large language models. Each brings its own strengths to different protein analysis tasks.
The team has made everything open source and available on GitHub. Their pre-trained models live on HuggingFace, ready for researchers to download and use. They've eliminated the need for redundant training, making model deployment faster and more efficient.
Why this matters:
- DeepProtein democratizes AI-powered protein research. What once required expertise in both biology and deep learning now needs just a basic understanding of command-line interfaces
- The comprehensive benchmarking across different AI architectures gives researchers clear guidance on which tools work best for specific protein analysis tasks. No more guessing games or trial and error
Read on, my dear:
- Deep Protein Models: All DeepProt-T5 models
 IMPLICATOR
IMPLICATOR



